Rfam is a collection of multiple sequence alignments and covariance models covering many common non-coding RNA families. The main use of Rfam is as a source of RNA multiple alignments with consensus secondary structure annotation in a consistent format. In conjunction with the Infernal software package, Rfam covariance models (CMs) can be used to search genomes or other DNA sequence databases for homologs to known structural RNA families.
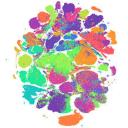
...continue to readCZ CELLxGENE from The Chan Zuckerberg Initiative, is a free-to-use online data portal collecting free and open-source tools for finding, querying, analysing, downloading and publishing single-cell data. As of April 2024, it includes some 85 million single cells 248 and 1,317 data sets covering 844 cell types. Most of the data represent single-cell RNA sequencing information from healthy human tissues, but non-human and cell-line data, as well as molecular-profiling data obtained using spatial tr
...continue to readAREsite is an online resource for the detailed investigation of AU-rich elements (ARE) in vertebrate mRNA 30-untranslated regions (UTRs. In its current version AREsite uses Ensembl release 56 as data basis (Gruber et al., Nucleic Acid Res 39:D66, 2011).
...continue to readSILVA is a comprehensive on-line resource for quality checked and aligned ribosomal RNA sequence data, free for academic use. SILVA provides comprehensive, quality checked and regularly updated datasets of aligned small (16S/18S, SSU) and large subunit (23S/28S, LSU) ribosomal RNA (rRNA) sequences for all three domains of life (Bacteria, Archaea and Eukarya).
...continue to readASPicDB is a database designed to provide access to reliable annotations of the alternative splicing pattern of human genes, obtained by ASPic algorithm, and to the functional annotation of predicted isoforms (Martelli et al., Nucleic Acids Res 39:D80, 2011)
...continue to readNONCODE is a database of all kinds of noncoding RNAs (except tRNAs and rRNAs).
...continue to readPage: 1